Code
getwd()[1] "/Users/leej/Documents/codex-managed/projects/research-hub/analysis"Grip Strength Patterns Across Athletic and Population Cohorts
Exploratory frequentist analysis comparing dominant-hand grip strength across SG25 athletes, NHANES exercisers, and NHANES non-exercisers, with within-group contrasts by self-reported high blood pressure (HBP).
library(dplyr)
library(tidyr)
prepared <- combined %>%
mutate(
meets_guidelines = case_when(
meets_ACSM_guidelines %in% c(TRUE, 1, "Yes", "Y") ~ TRUE,
meets_ACSM_guidelines %in% c(FALSE, 0, "No", "N") ~ FALSE,
TRUE ~ NA
),
activity_group = case_when(
dataset == "SG25" ~ "SG_Athletes",
dataset == "NHANES" & meets_guidelines %in% TRUE ~ "NHANES_Exercisers",
dataset == "NHANES" & meets_guidelines %in% FALSE ~ "NHANES_NonExercisers",
TRUE ~ NA_character_
),
hbp_status = if_else(HBP %in% c(TRUE, 1, "Yes", "Y"), "HBP_Yes", "HBP_No")
) %>%
filter(!is.na(activity_group), is.finite(grip_dom))
cohorts <- prepared
dplyr::count(cohorts, activity_group, hbp_status)# A tibble: 6 × 3
activity_group hbp_status n
<chr> <chr> <int>
1 NHANES_Exercisers HBP_No 1136
2 NHANES_Exercisers HBP_Yes 1101
3 NHANES_NonExercisers HBP_No 835
4 NHANES_NonExercisers HBP_Yes 1286
5 SG_Athletes HBP_No 78
6 SG_Athletes HBP_Yes 37library(gt)
group_levels <- c("SG_Athletes", "NHANES_Exercisers", "NHANES_NonExercisers")
grip_summary_overall <- cohorts %>%
mutate(activity_group = factor(activity_group, levels = group_levels)) %>%
group_by(activity_group) %>%
summarise(
n = n(),
mean_grip = mean(grip_dom, na.rm = TRUE),
sd_grip = sd(grip_dom, na.rm = TRUE),
.groups = "drop"
) %>%
mutate(summary = sprintf("%.2f ± %.2f (n=%d)", mean_grip, sd_grip, n))
grip_summary_overall# A tibble: 3 × 5
activity_group n mean_grip sd_grip summary
<fct> <int> <dbl> <dbl> <chr>
1 SG_Athletes 115 38.9 11.7 38.89 ± 11.74 (n=115)
2 NHANES_Exercisers 2237 35.1 10.5 35.07 ± 10.48 (n=2237)
3 NHANES_NonExercisers 2121 30.7 10.3 30.73 ± 10.31 (n=2121)| Dominant-Hand Grip Strength by Activity Group | ||||
| Mean ± SD (n) across SG athletes and NHANES exerciser/non-exerciser cohorts | ||||
| activity_group | n | mean_grip | sd_grip | summary |
|---|---|---|---|---|
| SG_Athletes | 115 | 38.88870 | 11.74300 | 38.89 ± 11.74 (n=115) |
| NHANES_Exercisers | 2237 | 35.06737 | 10.48217 | 35.07 ± 10.48 (n=2237) |
| NHANES_NonExercisers | 2121 | 30.73442 | 10.31196 | 30.73 ± 10.31 (n=2121) |
grip_summary_within <- cohorts %>%
filter(!is.na(hbp_status)) %>%
mutate(activity_group = factor(activity_group, levels = group_levels)) %>%
group_by(activity_group, hbp_status) %>%
summarise(
n = n(),
mean_grip = mean(grip_dom, na.rm = TRUE),
sd_grip = sd(grip_dom, na.rm = TRUE),
.groups = "drop"
) %>%
mutate(
summary = sprintf("%.2f ± %.2f (n=%d)", mean_grip, sd_grip, n)
) %>%
select(activity_group, hbp_status, summary) %>%
pivot_wider(
names_from = hbp_status,
values_from = summary,
values_fill = "—"
) %>%
arrange(activity_group)
grip_summary_within# A tibble: 3 × 3
activity_group HBP_No HBP_Yes
<fct> <chr> <chr>
1 SG_Athletes 39.07 ± 11.78 (n=78) 38.51 ± 11.81 (n=37)
2 NHANES_Exercisers 35.28 ± 10.30 (n=1136) 34.85 ± 10.66 (n=1101)
3 NHANES_NonExercisers 31.86 ± 10.03 (n=835) 30.00 ± 10.43 (n=1286)| Dominant-Hand Grip Strength by Activity Group and HBP Status | ||
| Mean ± SD (n) stratified by hypertension history | ||
| activity_group | HBP_No | HBP_Yes |
|---|---|---|
| SG_Athletes | 39.07 ± 11.78 (n=78) | 38.51 ± 11.81 (n=37) |
| NHANES_Exercisers | 35.28 ± 10.30 (n=1136) | 34.85 ± 10.66 (n=1101) |
| NHANES_NonExercisers | 31.86 ± 10.03 (n=835) | 30.00 ± 10.43 (n=1286) |
library(broom)
library(purrr)
ttest_within_results <- cohorts %>%
filter(!is.na(hbp_status)) %>%
group_by(activity_group) %>%
summarise(
test = list(
if (n_distinct(hbp_status) == 2) {
tidy(t.test(grip_dom ~ hbp_status, conf.level = 0.95))
} else {
tibble(
estimate = NA_real_,
statistic = NA_real_,
p.value = NA_real_,
conf.low = NA_real_,
conf.high = NA_real_
)
}
),
.groups = "drop"
) %>%
tidyr::unnest(test) %>%
mutate(activity_group = factor(activity_group, levels = group_levels))
ttest_within_results# A tibble: 3 × 11
activity_group estimate estimate1 estimate2 statistic p.value parameter
<fct> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 NHANES_Exercisers 0.424 35.3 34.9 0.955 3.40e-1 2225.
2 NHANES_NonExercisers 1.86 31.9 30.0 4.11 4.21e-5 1831.
3 SG_Athletes 0.557 39.1 38.5 0.237 8.14e-1 70.7
# ℹ 4 more variables: conf.low <dbl>, conf.high <dbl>, method <chr>,
# alternative <chr># A tibble: 2 × 6
term df sumsq meansq statistic p.value
<chr> <dbl> <dbl> <dbl> <dbl> <dbl>
1 activity_group 2 24380. 12190. 112. 3.70e-48
2 Residuals 4470 486836. 109. NA NA # A tibble: 4 × 7
# Groups: hbp_status [2]
hbp_status term df sumsq meansq statistic p.value
<fct> <chr> <dbl> <dbl> <dbl> <dbl> <dbl>
1 HBP_Yes activity_group 2 15384. 7692. 69.0 7.31e-30
2 HBP_Yes Residuals 2421 269952. 112. NA NA
3 HBP_No activity_group 2 7669. 3835. 36.5 2.69e-16
4 HBP_No Residuals 2046 215026. 105. NA NA # A tibble: 6 × 5
term estimate std.error statistic p.value
<chr> <dbl> <dbl> <dbl> <dbl>
1 (Intercept) 39.1 1.18 33.1 0
2 activity_groupNHANES_Exercisers -3.79 1.22 -3.11 0.0019
3 activity_groupNHANES_NonExercisers -7.21 1.23 -5.84 0
4 hbp_statusHBP_Yes -0.557 2.08 -0.268 0.789
5 activity_groupNHANES_Exercisers:hbp_stat… 0.134 2.13 0.0628 0.950
6 activity_groupNHANES_NonExercisers:hbp_s… -1.30 2.13 -0.611 0.541 library(ggplot2)
grip_plot <- cohorts %>%
filter(!is.na(hbp_status)) %>%
mutate(
activity_group = factor(activity_group, levels = group_levels),
hbp_status = factor(hbp_status, levels = c("HBP_No", "HBP_Yes"))
) %>%
ggplot(aes(x = activity_group, y = grip_dom, fill = hbp_status)) +
geom_violin(alpha = 0.4, trim = FALSE) +
geom_boxplot(width = 0.12, position = position_dodge(width = 0.9), outlier.shape = NA) +
stat_summary(fun = mean, geom = "point", shape = 21, size = 2.5, position = position_dodge(width = 0.9)) +
scale_fill_manual(values = c("HBP_No" = "#1b9e77", "HBP_Yes" = "#d95f02")) +
labs(
title = "Grip Strength Distributions by Activity Group and HBP History",
x = NULL,
y = "Dominant-hand grip (kg)",
fill = "HBP Status"
) +
theme_minimal()
grip_plot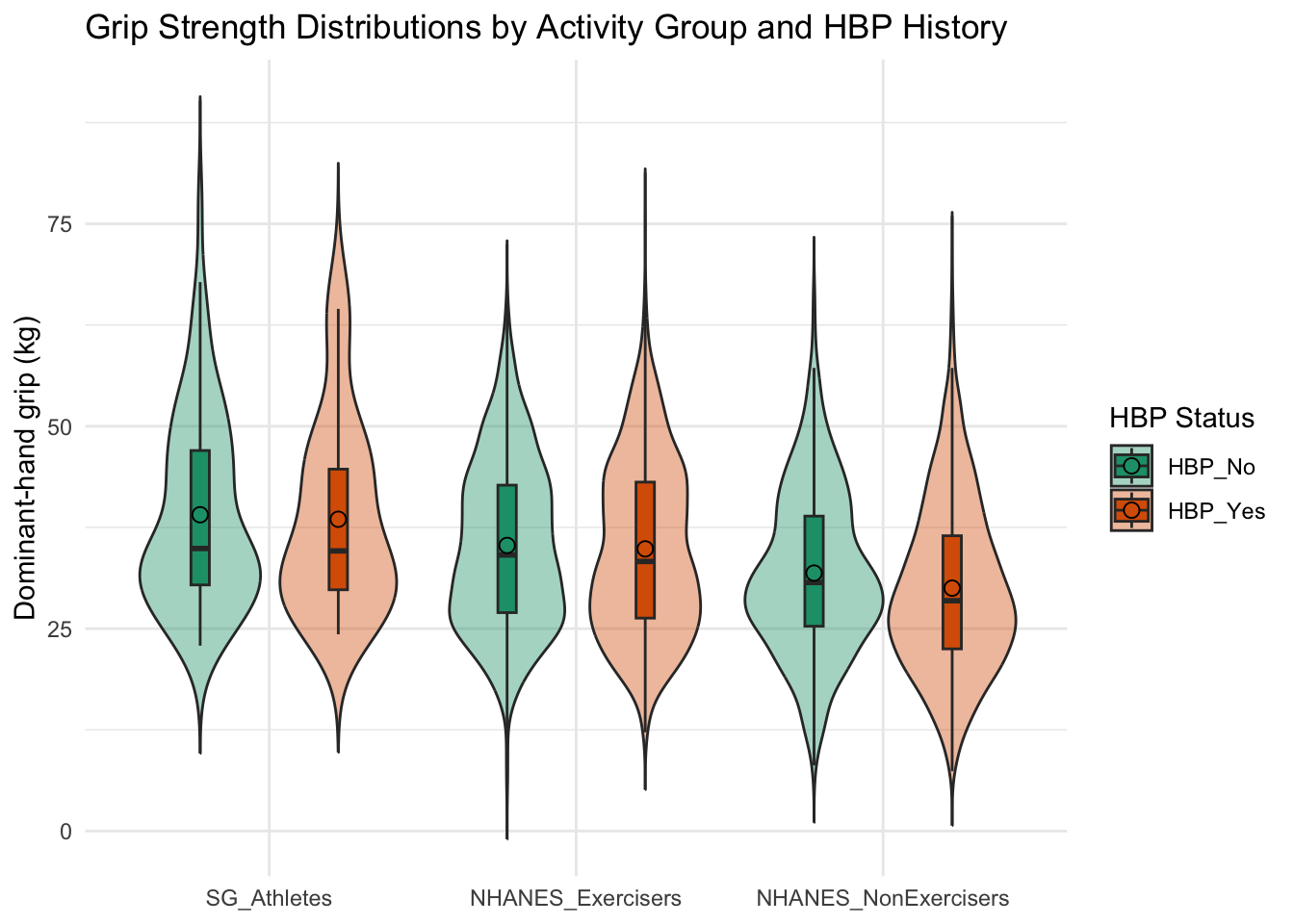
table_path <- here::here("analysis", "artifacts", "tables", "hbp-grip-summary.rds")
figure_path <- here::here("analysis", "artifacts", "figures", "hbp-grip-boxplot.png")
dir.create(dirname(table_path), showWarnings = FALSE, recursive = TRUE)
dir.create(dirname(figure_path), showWarnings = FALSE, recursive = TRUE)
saveRDS(
list(
overall = grip_summary_overall,
by_hbp = grip_summary_within
),
table_path
)
ggsave(figure_path, plot = grip_plot, width = 8, height = 5, dpi = 300)
list(table_rds = table_path, figure_png = figure_path)$table_rds
[1] "/Users/leej/Documents/codex-managed/projects/research-hub/analysis/artifacts/tables/hbp-grip-summary.rds"
$figure_png
[1] "/Users/leej/Documents/codex-managed/projects/research-hub/analysis/artifacts/figures/hbp-grip-boxplot.png"combined_NHANES_SG25_activity_harmonized.rds to capture updated cohorts.